In a paper presented at the International Conference on Machine Learning (ICML), researchers at the Massachusetts Institute of Technology developed a geometric deep learning model called EquiBind. It is 1,200 times faster than its predecessor and successfully binds drug molecules to proteins.
The current process for finding promising drug candidate molecules looks like this: a candidate sampling is combined with techniques such as scoring, ranking and fine-tuning, to get the best "match" between the ligand and the protein.
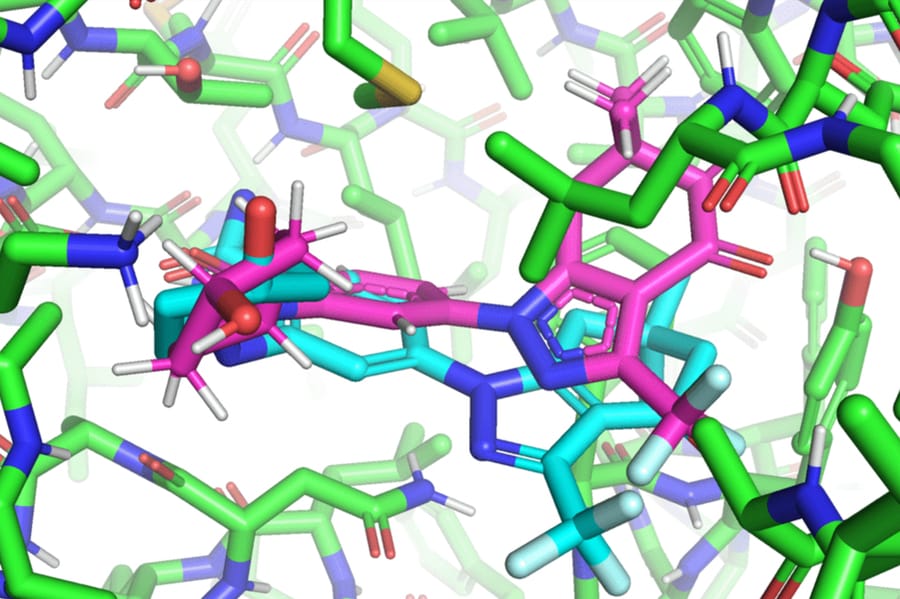
Unlike most models, which require multiple attempts to find a favorable ligand position in the protein, EquiBind already has built-in geometric reasoning that helps the model make better predictions when confronted with new, unseen data.
The EquiBind model has been tested on existing drugs and proteins used to treat lung cancer, leukemia, and gastrointestinal tumors. While most traditional docking methods have failed to successfully bind ligands that act on these proteins, EquiBind has succeeded.





Comments